goals and approach
The Trapnell Lab mission is to understand how the genome encodes the program of development and the role this program plays in disease. It develops and applies genomic tools, especially single-cell sequencing, to a variety of in vitro and animal model systems.
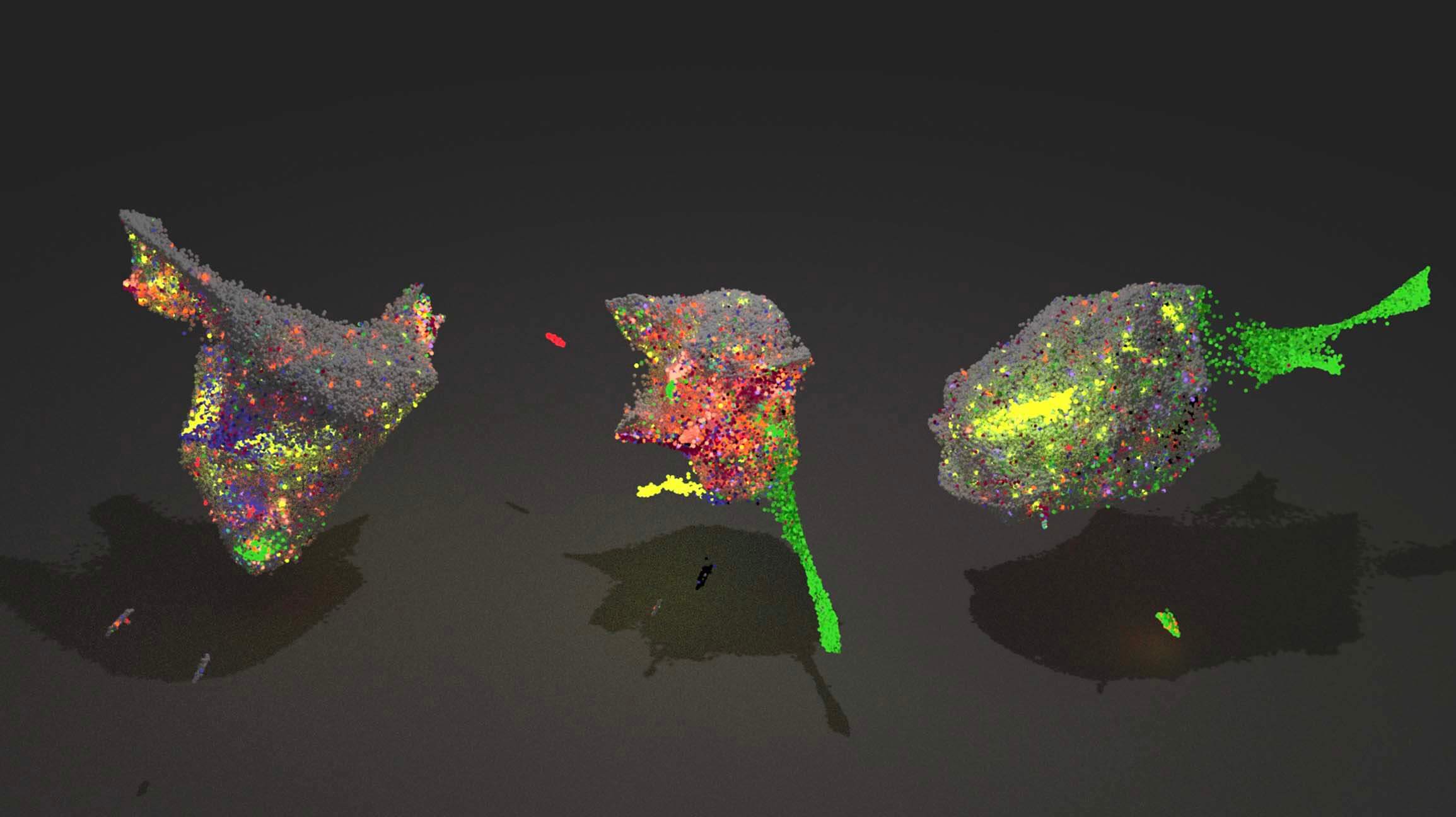
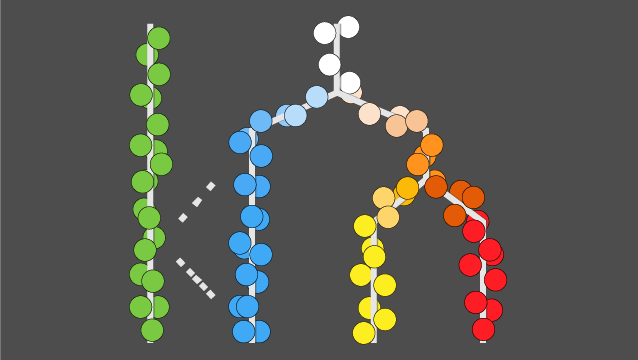
The lab’s research strategy is to develop new algorithms and measurements technologies that will enable the scientific community to quantitatively model how every gene is regulated in every cell type found in an animal. Lab members draw from computer science, statistics, and molecular cell biology to jointly develop both new assays and algorithms that fully exploit their capabilities. The scientists are using their technologies to understand the genetic control of plasticity in zebrafish and other key model systems of development.
Lead Investigator:
Dr. Cole Trapnell
Cole Trapnell is a Professor of Genome Sciences at the University of Washington. He received his Ph.D in Computer Science at the University of Maryland and has formal training in both computational and experimental biology. As a graduate student with Steven Salzberg (of Johns Hopkins University) and Lior Pachter (of the California Institute of Technology), , Dr. Trapnell wrote “TopHat” and “Cufflinks”, two widely used tools for transcriptome sequencing (RNA-seq) analysis. He also co-developed (with Johns Hopkins University’s Ben Langmead) the ultrafast short read alignment program “Bowtie.”
As a postdoctoral fellow, Dr. Trapnell augmented his computational work with experimental training focused on analysis of cell differentiation, and developed “single-cell trajectory analysis,” an approach for studying cell differentiation using single-cell RNA-Seq. Dr. Trapnell’s lab has introduced computational tools extracting biological insights from single-cell data, such as “Monocle,” as well as experimental workflows for ultra-scalable single-cell molecular profiling experiments. The lab has used these tools to explore pancreatic islet development, olfactory neurogenesis, and thyroid hormone-dependent pigmentation, and is currently applying them to dissect the genetic program of zebrafish embryonic development. Dr Trapnell is a recipient of ISCB Overton Prize (2018), NIH Director’s New Innovator Award, and the Alfred P. Sloan Foundation Research Fellowship (2015), and Damon Runyon Dale F. Frey Award for Breakthrough Scientist (2014).