
in this article
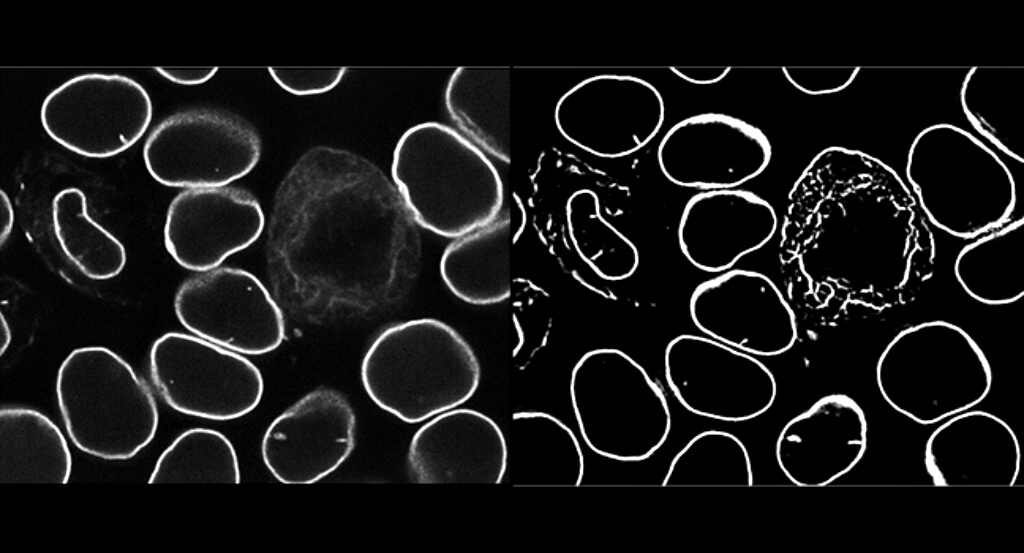
Biologists who want to find their way through the microscopic world inside a cell face a related challenge. To understand different components of the cell — and how they might change under important conditions, like growth or disease — you have to understand precisely where they are and what they look like. And that means knowing the exact location of those components’ edges, down to the pixel.
Enter the Allen Cell Structure Segmenter, a python-based, open-source toolkit incorporating machine learning techniques that does for cell biology what computer vision techniques have done for self-driving cars, broadly speaking. The tool helps scientists navigate complex images of cells by automatically capturing the boundaries of structures inside them.
The toolkit was developed by researchers at the Allen Institute for Cell Science, a division of the Allen Institute, as a way to probe 3D images in the Allen Cell Collection, a catalog of live human stem cell lines gene-edited so that different structures glow with fluorescent colors under the microscope. The collection includes 38 cell lines tagging 34 different cellular structures, with more coming down the pike. The researchers have collected an enormous number of images, and realized they needed an automated and accurate way to navigate the many different structures, whose shape in the fluorescent images varies from tiny dots to smooth boundaries to ruffled edges.
“We needed a way to access and define those structures for every single one of those cell lines,” said Susanne Rafelski, Ph.D., director of the Assay Development team at the Allen Institute for Cell Science, a division of the Allen Institute. “Given that every structure has its own set of challenges, and we’re going to have lots and lots of them, it gets really hard.”
And she knew that other cell biologists face similar challenges. Rafelski, along with Allen Institute for Cell Science researcher Jianxu Chen, Ph.D., and the rest of the Assay Development team, created the Allen Cell Structure Segmenter and decided to make the toolkit available publicly so other scientists could use it.
Not every cell biology question needs such a rigorous approach. In many cases, biologists can find structures in cells just by looking at the images. But in other situations — especially for scientists working with 3D images of cells — more precision or more automation is needed.
Two approaches for the cell biology community
The Segmenter toolkit was rolled out in two phases. Last year, the team debuted the first part, which gathers existing computer vision algorithms into a menu. This “classic workflow” part of the toolkit presents recommendations for segmenting a given image through a look-up table of representative pictures from the Allen Cell Collection. Researchers can scan the table and find images that look similar to their own. The recommendations depend on patterns of fluorescence — say, tiny speckles or large diffuse circles — rather than the specific structure identity.
The newest part of the toolkit, debuted this spring on allencell.org, uses machine learning to identify structure boundaries in images when the classic approach isn’t sufficient for the problem. For example, many structures in the cell change dramatically during the different stages of cell division. Some structures might look like a smooth circle most of the time, but quickly shift into a series of bright speckles as the cell divides. The machine learning part of the Segmenter toolkit can, with some straight-forward user input, tackle that problem.
“The classic workflow can work without the machine learning, and the machine learning can work without the classic,” Rafelski said. “We use them together and it’s really powerful.”
Github repositories for both parts of the Allen Cell Structure Segmenter are available on allencell.org, with a video tutorial available for the classic segmentation workflow. The team is working on additional tutorials and expansions to the tool as they receive community feedback.
“This is a nice resource to give back to the community,” said Derek Thirstrup, an image analyst at the Allen Institute for Cell Science who helped test the toolkit. “It’s really lowering the entry barrier to let cell biologists be more quantitative in their results.”
Citations
about the allen institute
The Allen Institute is an independent, 501(c)(3) nonprofit research organization founded by philanthropist and visionary, the late Paul G. Allen. The Allen Institute is dedicated to answering some of the biggest questions in bioscience and accelerating research worldwide. The Institute is a recognized leader in large-scale research with a commitment to an open science model. For more information, visit alleninstitute.org.